
histogram|-H = histogram computation level: (-H alone implies -H1)ġ - basic bucketsize/bucketcount histogram dataĢ - bucket-based scores to tils hitfilter|-h = minimum number of hits to be outputted as an anchor probing = 0 - linear, 1 - quadratic (default) MSetHash - memory exorbanant, almost pointless. OpenHash - open sub-word packing of hashbitsĮcoHash - chained sub-word packing of hashbits (default)ĪrrayHash - malloc/realloc (fast but fragmenty)
hashtype|-t = select hash table data structure to use: hashbits|-b = use n bit hashes (for n's of 1 to WORDSIZE. quickhash|-q = specify a hashing function:ġ - don't pack bits to make hash (use first word only)Ģ - naively use the first hashbits worth of patternģ - adaptivevely find a good hash (default)

name|-n = alignment name (default: test)
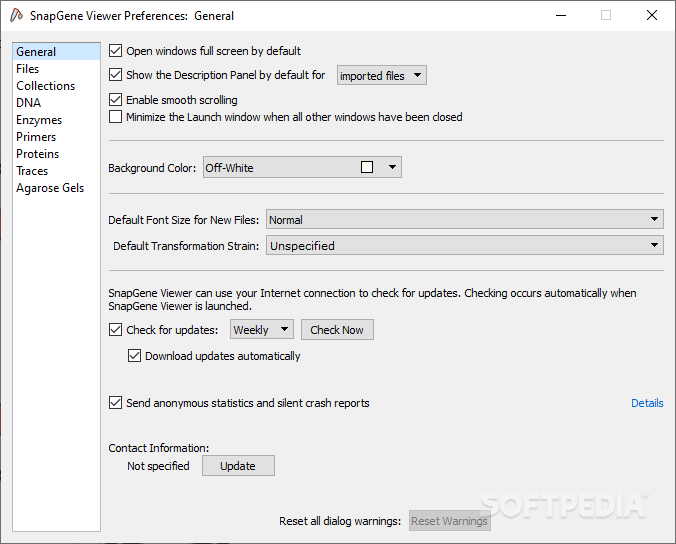
directory|-d = output directory (default: output) Using the format automatically generates a $ docker run -rm -it biocontainers/murasaki:v1.68.6-8b1- deb_cv1 murasaki -h > docker run -rm -it biocontainers/murasaki:v1.68.6-8b1- deb_cv1 murasaki -h MUMmer4: A fast and versatile genome alignment system Stefan Kurtz*, Adam Phillippy†, Arthur L Delcher†, Michael Smoot†‡, Martin Shumway†, Corina Antonescu† and Steven L Salzberg†
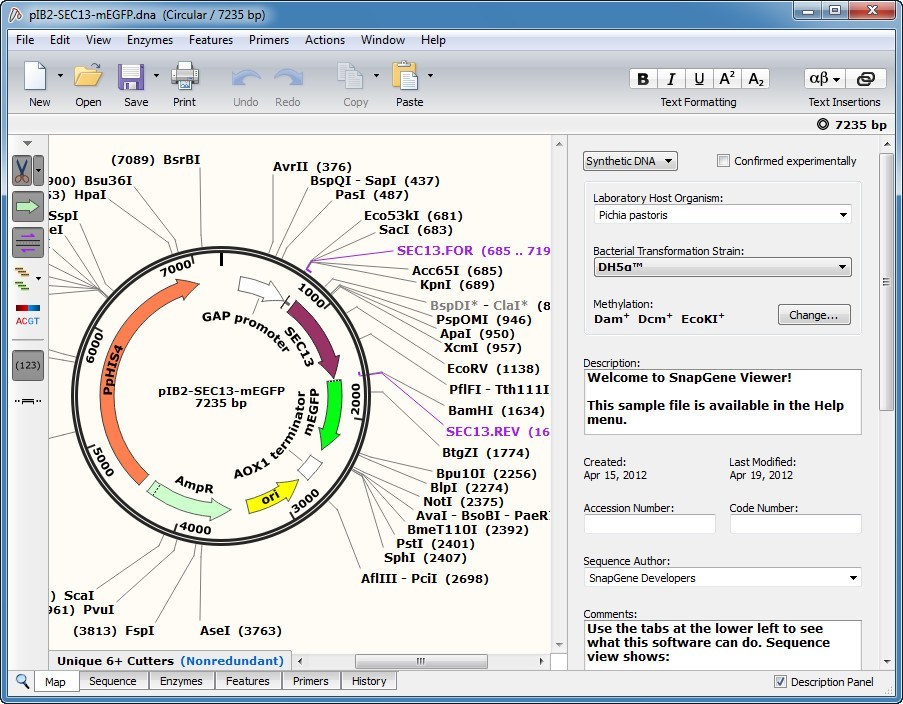
#Snapgene viewer 32 bit software
3、 Versatile and open software for comparing large genomes 2、 Fast algorithms for large-scale genome alignment and comparison.ĭelcher AL1, Phillippy A, Carlton J, Salzberg SL. SnapGene software (from GSL Biotech available at ) SnapGene Viewer | Molecular Biology Software SnapGene Software Tutorial Videos | Aligning to a Reference DNA Sequence SnapGene | Software for everyday molecular biology

